|
The data show how the different architectures of chaperones result in distinct binding modes with non-native proteins that ultimately define the activity of the chaperone.ĭepartment of Biochemistry, Molecular Biology &Biophysics, University of Minnesota, Minneapolis, Minnesota 55455, USA. This unique complex architecture alters the kinetics of protein binding to SecB and confers strong antifolding activity on the chaperone. The protein may fold into its native structure on the chaperone surface or upon release. The multivalent binding mode results in proteins wrapping around SecB. The PapD-like superfamily of periplasmic chaperones directs the assembly of over 30 diverse adhesive surface organelles that mediate the attachment of many different pathogenic bacteria to host tissues, a critical early step in the development of disease ( 1, 2 ). The chaperone features a chaotropic pocket to bind and stabilize the protein, protecting it from premature hydrophobic collapse into a misfolded state and allowing it to explore the conformational space. SecB uses long hydrophobic grooves that run around its disk-like shape to recognize and bind to multiple hydrophobic segments across the length of non-native proteins. Here we report the solution structure of SecB, a chaperone that exhibits strong antifolding activity, in complex with alkaline phosphatase and maltose-binding protein captured in their unfolded states. that their three-dimensional structure is almost always wrong by that point. Different chaperones often exert distinct effects, such as acceleration or delay of folding, on client proteins via mechanisms that are poorly understood. But the aggregated, misfolded proteins are also insoluble, which can make. Molecular chaperones act on non-native proteins in the cell to prevent their aggregation, premature folding or misfolding. Diversity, Equity, Inclusion, and Access.Citation, Usage, Privacy Policies, Logo.Biologically Interesting Molecule Reference Dictionary (BIRD) In 1917 the German chemist Hermann Staudinger proposed that organic molecules such as proteins were organized in polymers, giant molecules made of small-molecule constituents linked together by.Note that because of processes such as the post-translational modifications to proteins we still need protein sequencing and I believe that we currently rely too heavily on DNA sequencing. This is because it is now much easier to sequence DNA. It is clear that molecular chaperones assist with the folding of newly synthesized proteins and correct protein misfolding. Instead, since it has been worked out (mostly) how DNA codes for protein, we usually infer the protein sequence from the DNA sequence. Proteins are not usually linear, but fold naturally into organized structures called native structures. The structure of the AB fragment is determined and stabilized purely by the interactions.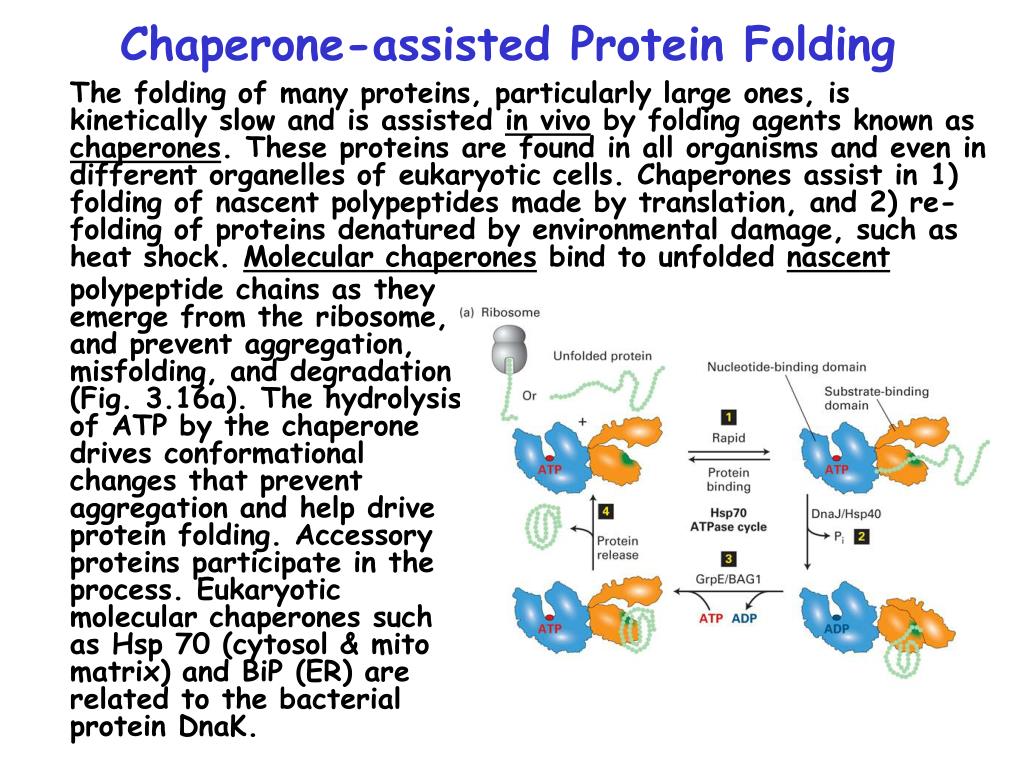 
Under physiological conditions, the in vitro folding of Tetrahymena ribozyme by the RNA chaperone CYT-19 behaves paradoxically increasing the chaperone concentration reduces the yield of native ribozymes. obviously does not determine the native three-dimen-. First is the fact that proteins are only marginally stable. Molecular chaperones facilitate the folding of proteins and RNA in vivo. However, it is now relatively rare to directly determine protein sequence! differences: nascent proteins acquire their native structure co- and post-translationally, with. The very first protein sequence (bovine insulin) was determined by Fredrick Sanger in 1951-2 (note that this was more than a decade before the first nucleotide sequence). Chaperones exist in all cellular compartments and interact with the polypeptide chain in order to allow the native three-dimensional conformation of the protein. There are many different techniques for directly determining protein sequences - this wikipedia article is a decent introduction: Proteins that help other proteins during their folding and are involved in protein homeostasis are called molecular chaperones. 
There are also methods that have been developed to remove amino acids one at a time.īy combining theses techniques it is possible to directly determine protein sequences. Chaperones can avoid the conformational change to beta sheet. This is a great question, but actually quite complicated so I'm not going to try to give a complete answer - I have given some useful links below if you wish to learn more.Įach amino acid has unique chemical properties that can be used to tell them apart. Proteins that have problems achieving their native configuration are helped by chaperones to fold properly, using energy from ATP.
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
